Master Univisual
Understand how Univisual's settings shape your analysis results.
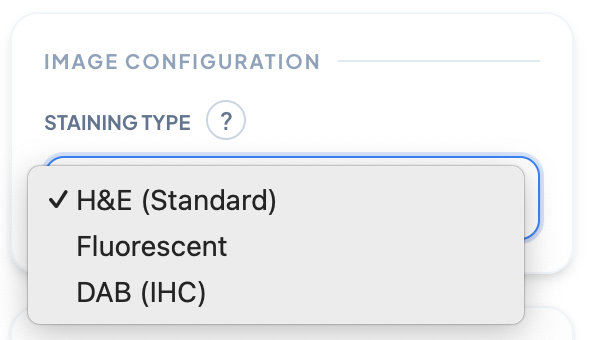
Staining Type
Select the type of biological staining used in your microscopy image. Different stains (H&E, Fluorescent, DAB/IHC) highlight different cellular structures, affecting how the algorithm processes and analyzes your sample.
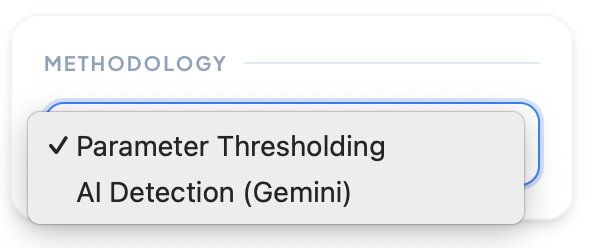
Methodology
Choose your detection method: Parameter Thresholding for manual fine-tuning of detection parameters, or AI Detection (Gemini) for automated cell identification using advanced machine learning algorithms.
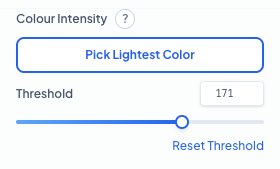
Colour Intensity Parameters
Fine-tune the detection sensitivity by setting the brightness threshold. Use 'Pick Lightest Color' to automatically sample the peak intensity from your cell of interest. **Threshold:** This defines the maximum brightness allowed for a pixel to be considered part of a cell. Lowering this value captures fainter signals but may increase noise, while raising it focuses only on the most intensely stained regions.
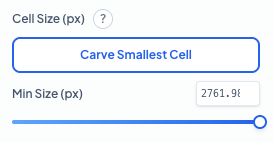
Cell Size Parameters
Filter out background noise and artifacts by setting a minimum cell size. Use 'Carve Smallest Cell' to interactively define the smallest object you want the algorithm to count. **Min Size (px):** Objects smaller than this pixel area will be excluded from the final count. This is essential for ensuring that only complete, valid cells are quantified while ignoring small debris or staining artifacts.
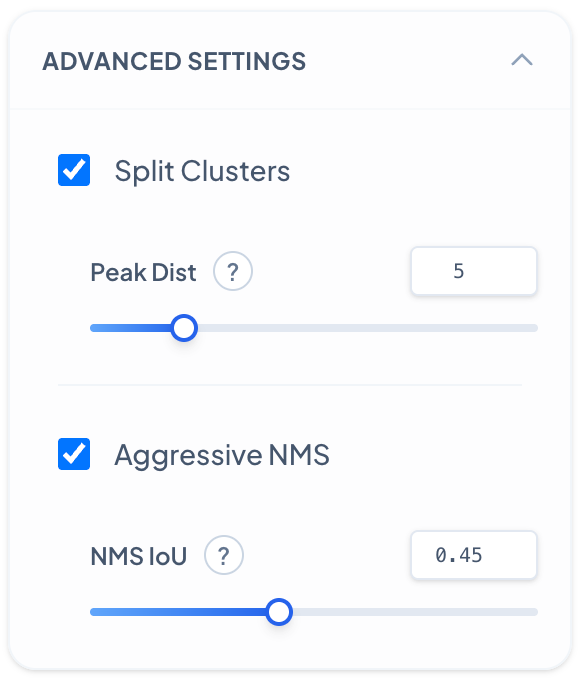
Advanced Settings - Peak Distance
Controls how the algorithm separates touching or overlapping cell clusters based on local intensity peaks. **Peak Distance:** Decreasing results in more aggressive splitting of cell clusters, detecting more individual cells within dense regions but may over-segment single cells. Increase when cells are well-separated to avoid false splits and maintain cluster integrity.

Advanced Settings - NMS IoU
Non-Maximum Suppression (NMS) IoU threshold controls how duplicate detections are filtered. **NMS IoU Threshold:** Decreasing makes the algorithm more aggressive at removing overlapping detections, reducing duplicates but may eliminate valid closely-spaced cells. Increase to preserve more detections in dense regions where cells naturally overlap.
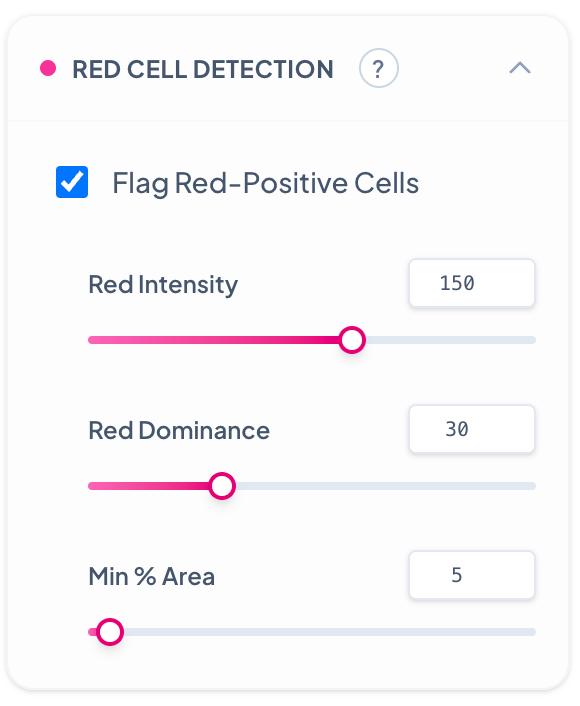
Red Cell Detection
Enables detection and flagging of cells showing positive signal in the red channel, ideal for immunofluorescence or lipid phenotyping. **Red Intensity:** Decreasing detects cells with weak red fluorescence signals but may include auto-fluorescence and background. Increase to target only cells with strong red markers. **Red Dominance:** Decreasing flags cells with any red signal presence regardless of other channels. Increase to require red signal to be dominant over other color channels. **Min % Area:** Decreasing requires only small portions of a cell to be red-positive for flagging. Increase to require larger red-positive areas within cells, ensuring more confident red cell identification.